|
| 1 | + |
| 2 | + |
| 3 | +# AlphaGenome |
| 4 | +[**Overview**](#overview) | [**Use Cases**](#use-cases) | [**Documentation**](#documentation) | [**Pricing**](#pricing) | [**Quick start**](#quick-start) |
| 5 | + |
| 6 | +## Overview |
| 7 | +**Disclaimer:** *Experimental*. |
| 8 | + |
| 9 | +*The AlphaGenome Private Preview is a "Pre-GA Offering" subject to the "Pre-GA |
| 10 | +Offerings Terms" in the General Service Terms section of the Google Cloud |
| 11 | +[Service Specific Terms](https://cloud.google.com/terms/service-terms). It is |
| 12 | +also a “Generative AI Preview Product” as defined in and subject to the |
| 13 | +[Additional Terms for Generative AI Preview Products](https://cloud.google.com/trustedtester/aitos?e=48754805&hl=en). |
| 14 | +Pre-GA products are available "as is" and might have limited support. For more |
| 15 | +information, see the [launch stage](https://cloud.google.com/products?e=48754805#product-launch-stages) |
| 16 | +descriptions.* |
| 17 | + |
| 18 | +Access to the AlphaGenome model capabilities requires application and approval. |
| 19 | +Users must be added to an allowlist to use the service. |
| 20 | +If you are interested in applying to the program, **Request Access** above. |
| 21 | + |
| 22 | + |
| 23 | + |
| 24 | +AlphaGenome is Google DeepMind’s unifying model for deciphering the regulatory |
| 25 | +code within DNA sequences. |
| 26 | + |
| 27 | +AlphaGenome offers multimodal predictions, encompassing diverse functional |
| 28 | +outputs such as gene expression, splicing patterns, chromatin features, and |
| 29 | +contact maps (see diagram below). The model analyzes DNA sequences of up to 1 |
| 30 | +million base pairs in length and can deliver predictions at single base-pair |
| 31 | +resolution for most outputs. |
| 32 | + |
| 33 | +Training data was sourced from large public consortia including |
| 34 | +[ENCODE](http://encodeproject.org/), [GTEx](https://www.gtexportal.org/), |
| 35 | +[4D Nucleome](https://4dnucleome.org/) and |
| 36 | +[FANTOM5](https://fantom.gsc.riken.jp/5/), which experimentally measured these |
| 37 | +properties covering important modalities of gene regulation across hundreds of |
| 38 | +human and mouse cell types and tissues. |
| 39 | + |
| 40 | +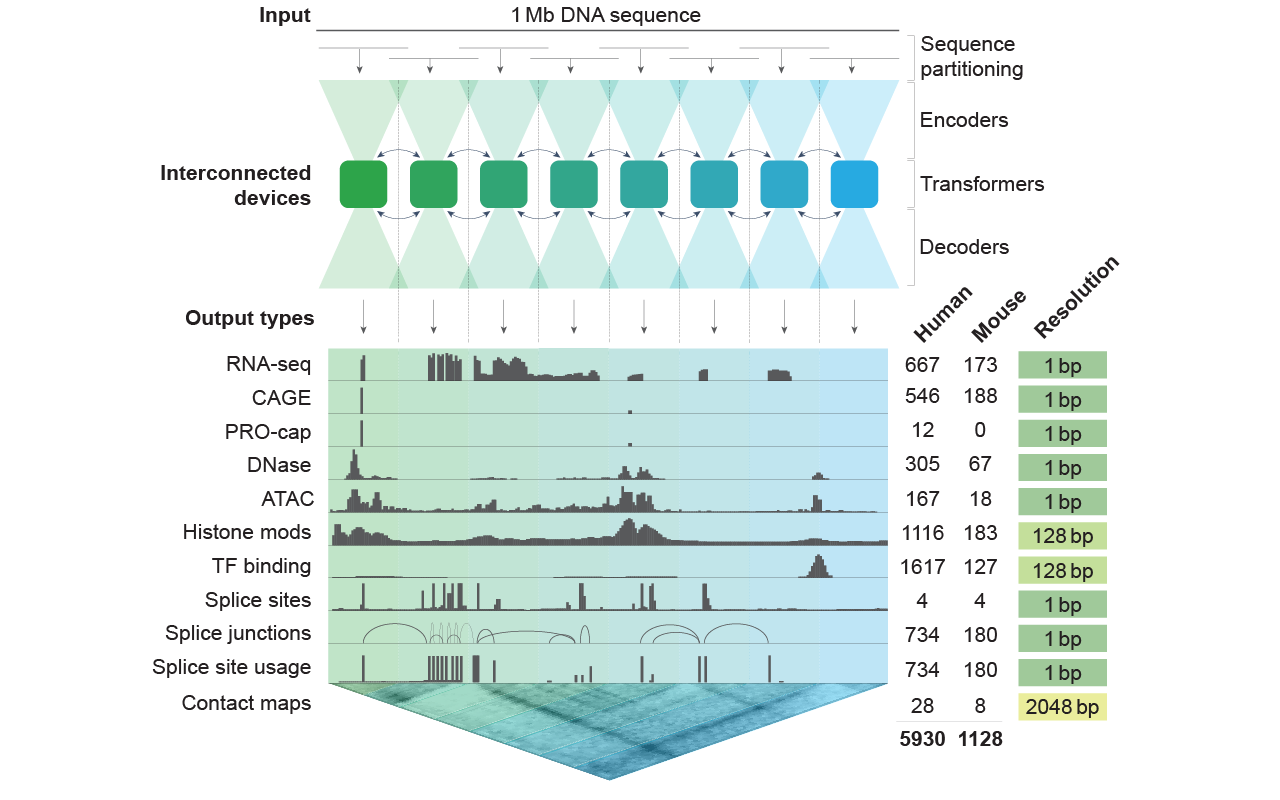 |
| 41 | + |
| 42 | +## Use Cases |
| 43 | +* **Predict outputs for a DNA sequence:** AlphaGenome is a model that makes |
| 44 | + predictions from DNA sequences. AlphaGenome predicts multiple 'tracks' per |
| 45 | + output type, covering a wide variety of tissues and cell-types. |
| 46 | + |
| 47 | +* **Open-vocabulary object retrieval:** AlphaGenome can make predictions for |
| 48 | + a human reference genome sequence specified by a genomic interval. |
| 49 | + For example, let's predict RNA-seq for tissue 'Right liver lobe' in a 1MB |
| 50 | + region of Chromosome 19 around the gene CYP2B6, which encodes an enzyme |
| 51 | + involved in drug metabolism, and is primarily expressed in the liver. |
| 52 | + |
| 53 | +* **Predict variant effects:** AlphaGenome can predict the effect of a |
| 54 | + variant on a specific output type and tissue by making predictions for the |
| 55 | + reference (REF) and alternative (ALT) allele sequences. |
| 56 | + |
| 57 | +* **Scoring the effect of a genetic variant:** AlphaGenome can score the |
| 58 | + effect of a genetic variant by making predictions for the REF and ALT |
| 59 | + sequences and aggregating the track signal. To highlight which regions in a |
| 60 | + DNA sequence are functionally important for a final variant prediction, |
| 61 | + AlphaGenome can help you to perform an in silico mutagenesis (ISM) analysis |
| 62 | + by scoring all possible single nucleotide variants in a specific interval. |
| 63 | + |
| 64 | +* **Human and mouse predictions:** AlphaGenome can generate predictions for |
| 65 | + both humans and mouse. |
| 66 | + |
| 67 | +## Documentation |
| 68 | +This API provides access to AlphaGenome, Google DeepMind's unifying model for |
| 69 | +deciphering the regulatory code within DNA sequences. AlphaGenome offers |
| 70 | +multimodal predictions, encompassing diverse functional outputs including gene |
| 71 | +expression, splicing patterns, chromatin features, and contact maps (see diagram |
| 72 | +below). The model analyzes up to 1 million base pairs of DNA sequence and can |
| 73 | +deliver predictions at single base-pair resolution for most modalities. |
| 74 | +AlphaGenome achieves state-of-the-art performance across a range of genomic |
| 75 | +prediction benchmarks, including diverse variant effect prediction tasks. |
| 76 | + |
| 77 | +The Google Cloud API for AlphaGenome provides a way for Google Cloud customers |
| 78 | +to explore the AlphaGenome API for commercial use cases. This API is in private |
| 79 | +preview (Request Access above). Once allowlisted, customers can access the API |
| 80 | +directly or use the [colab](cloudai_alphagenome_vai_quickstart.ipynb). |
| 81 | + |
| 82 | +### Acknowledgements |
| 83 | + |
| 84 | +*Avsec, Ž., Latysheva, N., Cheng, J., Novati, G., Taylor, K. R., Ward, T., ... Kohli, P. (2025). AlphaGenome: advancing regulatory variant effect prediction with a unified DNA sequence model. bioRxiv.* [https://doi.org/10.1101/2025.06.25.661532](https://doi.org/10.1101/2025.06.25.661532) |
| 85 | + |
| 86 | +### Contact |
| 87 | +If you have any questions on using these models on Google Cloud please contact: |
| 88 | +[alphagenome-cloud-external@google.com](mailto:alphagenome-cloud-external@google.com) or join the community [Discourse](https://www.alphagenomecommunity.com/) for more generic questions on AlphaGenome. |
| 89 | + |
| 90 | +### Links |
| 91 | + |
| 92 | +* Read our [paper](https://doi.org/10.1101/2025.06.25.661532) |
| 93 | +* Read our [blog post](https://deepmind.google/discover/blog/alphagenome-ai-for-better-understanding-the-genome) |
| 94 | +* Join the [community](https://www.alphagenomecommunity.com/) |
| 95 | +* Check out the [AlphaGenome 101 Video](https://youtu.be/Xbvloe13nak) |
| 96 | + |
| 97 | +## Pricing |
| 98 | +Access to AlphaGenome on Vertex AI is currently restricted. |
| 99 | +To utilize these models via this service: |
| 100 | + |
| 101 | +* You must **Request Access** using your Google contact. |
| 102 | +* Your application will be reviewed, and if approved, you will be **added to |
| 103 | + an allowlist**. |
| 104 | +* Only allowlisted users can access the API |
| 105 | +* **Pricing information** will be shared directly with users upon approval |
| 106 | + and placement on the allowlist. |
| 107 | + |
| 108 | +## Quick start |
| 109 | +The quickest way to get started with the AlphaGenome in Google Cloud Platform is to run [our example notebook](cloudai_alphagenome_vai_quickstart.ipynb) in [Google Colab](https://colab.research.google.com/). |
0 commit comments